Note
Click here to download the full example code
Plotting a joint histogram¶
In this recipe, we will be creating a joint histogram of PM2.5 mass concentration and 2-metre temperature.
Import cf-python, numpy and matplotlib.pyplot:
import matplotlib.pyplot as plt
import numpy as np
import cf
Read the field constructs using read function:
[<CF Field: long_name=Total Aerosol Optical Depth at 550nm(long_name=time(6), long_name=latitude(241), long_name=longitude(480)) ~>,
<CF Field: mass_concentration_of_pm10_ambient_aerosol_particles_in_air(long_name=time(6), long_name=latitude(241), long_name=longitude(480)) kg m**-3>,
<CF Field: mass_concentration_of_pm2p5_ambient_aerosol_particles_in_air(long_name=time(6), long_name=latitude(241), long_name=longitude(480)) kg m**-3>,
<CF Field: long_name=2 metre temperature(long_name=time(6), long_name=latitude(241), long_name=longitude(480)) K>]
3. Select the PM2.5 mass concentration and 2-metre temperature fields by index and print the description to show properties of all constructs:
pm25_field = f[2]
pm25_field.dump()
---------------------------------------------------------------------------------
Field: mass_concentration_of_pm2p5_ambient_aerosol_particles_in_air (ncvar%pm2p5)
---------------------------------------------------------------------------------
Conventions = 'CF-1.6'
_FillValue = -32767
history = '2023-05-02 11:17:49 GMT by grib_to_netcdf-2.25.1: /opt/ecmwf/mars-
client/bin/grib_to_netcdf.bin -S param -o /cache/tmp/4e68aba0-50a9-
4f06-947c-9fc7444b7838-adaptor.mars.internal-1683026269.508034-
23606-13-tmp.nc /cache/tmp/4e68aba0-50a9-4f06-947c-9fc7444b7838-
adaptor.mars.internal-1683026265.2095394-23606-10-tmp.grib'
long_name = 'Particulate matter d <= 2.5 um'
missing_value = -32767
standard_name = 'mass_concentration_of_pm2p5_ambient_aerosol_particles_in_air'
units = 'kg m**-3'
Data(long_name=time(6), long_name=latitude(241), long_name=longitude(480)) = [[[1.5032209791448226e-10, ..., 1.1372701346394527e-10]]] kg m**-3
Domain Axis: long_name=latitude(241)
Domain Axis: long_name=longitude(480)
Domain Axis: long_name=time(6)
Dimension coordinate: long_name=time
calendar = 'gregorian'
long_name = 'time'
units = 'hours since 1900-01-01 00:00:00.0'
Data(long_name=time(6)) = [2022-01-01 00:00:00, ..., 2022-06-01 00:00:00] gregorian
Dimension coordinate: long_name=latitude
long_name = 'latitude'
units = 'degrees_north'
Data(long_name=latitude(241)) = [90.0, ..., -90.0] degrees_north
Dimension coordinate: long_name=longitude
long_name = 'longitude'
units = 'degrees_east'
Data(long_name=longitude(480)) = [0.0, ..., 359.25] degrees_east
temp_field = f[3]
temp_field.dump()
------------------------------------------------
Field: long_name=2 metre temperature (ncvar%t2m)
------------------------------------------------
Conventions = 'CF-1.6'
_FillValue = -32767
history = '2023-05-02 11:17:49 GMT by grib_to_netcdf-2.25.1: /opt/ecmwf/mars-
client/bin/grib_to_netcdf.bin -S param -o /cache/tmp/4e68aba0-50a9-
4f06-947c-9fc7444b7838-adaptor.mars.internal-1683026269.508034-
23606-13-tmp.nc /cache/tmp/4e68aba0-50a9-4f06-947c-9fc7444b7838-
adaptor.mars.internal-1683026265.2095394-23606-10-tmp.grib'
long_name = '2 metre temperature'
missing_value = -32767
units = 'K'
Data(long_name=time(6), long_name=latitude(241), long_name=longitude(480)) = [[[247.0076398602821, ..., 226.25710500984027]]] K
Domain Axis: long_name=latitude(241)
Domain Axis: long_name=longitude(480)
Domain Axis: long_name=time(6)
Dimension coordinate: long_name=time
calendar = 'gregorian'
long_name = 'time'
units = 'hours since 1900-01-01 00:00:00.0'
Data(long_name=time(6)) = [2022-01-01 00:00:00, ..., 2022-06-01 00:00:00] gregorian
Dimension coordinate: long_name=latitude
long_name = 'latitude'
units = 'degrees_north'
Data(long_name=latitude(241)) = [90.0, ..., -90.0] degrees_north
Dimension coordinate: long_name=longitude
long_name = 'longitude'
units = 'degrees_east'
Data(long_name=longitude(480)) = [0.0, ..., 359.25] degrees_east
Convert the units to degree celsius for the temperature field:
temp_field.units = "degC"
temp_field.get_property("units")
'degC'
5. Digitize the PM2.5 mass concentration and 2-metre temperature fields.
This step counts the number of values in each of the 10 equally sized bins
spanning the range of the values. The 'return_bins=True' argument makes
sure that the calculated bins are also returned:
pm25_indices, pm25_bins = pm25_field.digitize(10, return_bins=True)
temp_indices, temp_bins = temp_field.digitize(10, return_bins=True)
Create a joint histogram of the digitized fields:
Get histogram data from the
joint_histogram:
Calculate bin centers for PM2.5 and temperature bins:
pm25_bin_centers = [(a + b) / 2 for a, b in pm25_bins]
temp_bin_centers = [(a + b) / 2 for a, b in temp_bins]
9. Create grids for PM2.5 and temperature bins using numpy.meshgrid:
10. Plot the joint histogram using matplotlib.pyplot.pcolormesh. Use cf.Field.unique to get
the unique data array values and show the bin boundaries as ticks on the
plot. 'style='plain' in matplotlib.axes.Axes.ticklabel_format
disables the scientific notation on the y-axis:
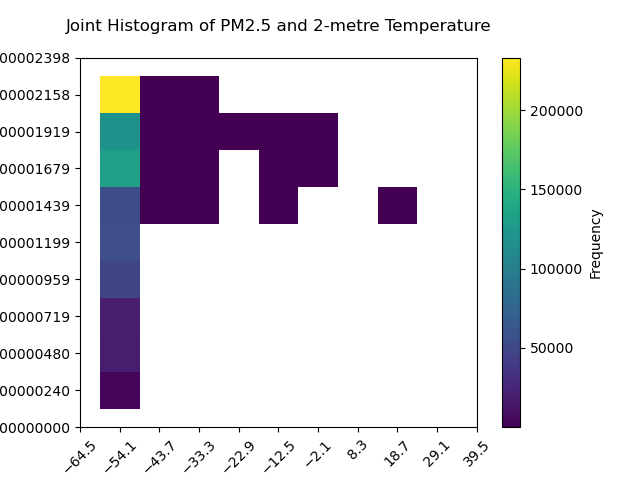
plt.pcolormesh(temp_grid, pm25_grid, histogram_data, cmap="viridis")
plt.xlabel("2-metre Temperature (degC)")
plt.ylabel("PM2.5 Mass Concentration (kg m**-3)")
plt.xticks(temp_bins.unique().array, rotation=45)
plt.yticks(pm25_bins.unique().array)
plt.colorbar(label="Frequency")
plt.title("Joint Histogram of PM2.5 and 2-metre Temperature", y=1.05)
plt.ticklabel_format(style="plain")
plt.show()
Total running time of the script: ( 0 minutes 4.643 seconds)